 Various extensions were added, including additional scriptingĬommands. The Chime web browser plug-in was based upon the RasMol source code. The current URL for full RasMol documentation Jmol supports the RasMol scripting language defined by RasMol. Supply feedback in case of bugs and/or unexpected behavior and/or 
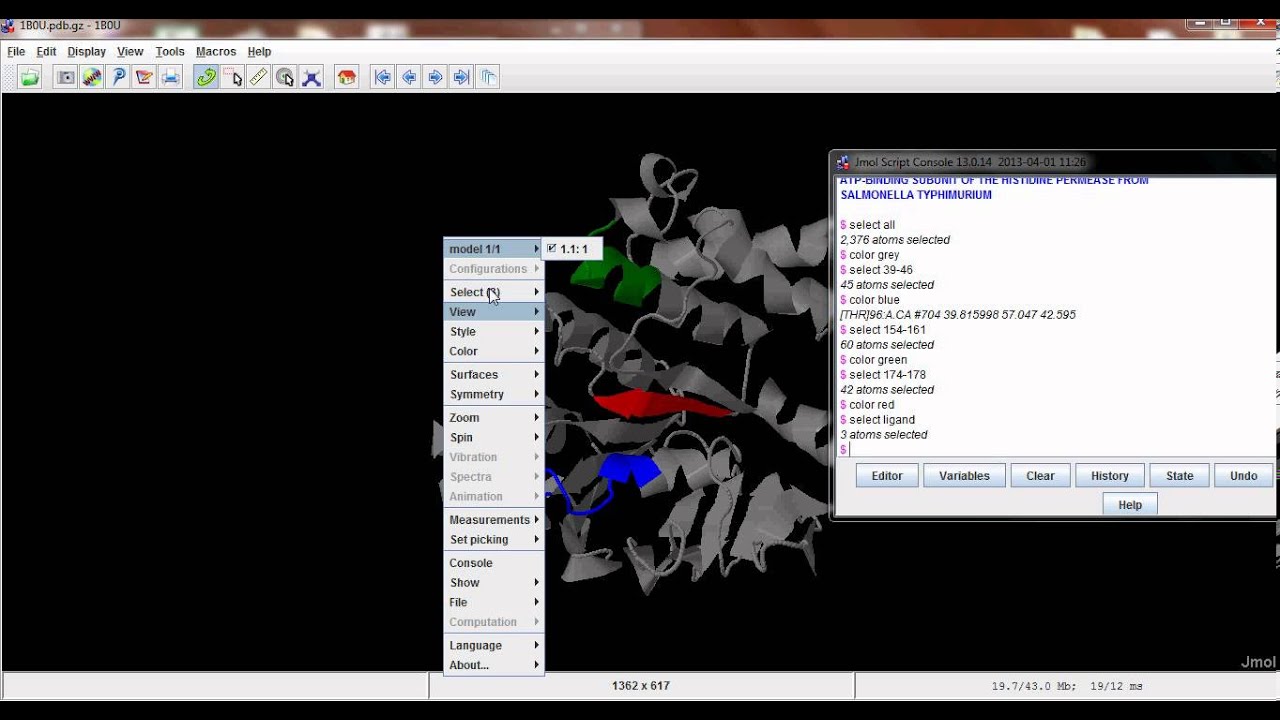
As of this writing it should be considered *alpha test* code. Open the molden.inp file with molden and write out a new file (select "write" in the moden menu).The scripting engine has been completely rewritten for Jmol ReleaseĦ. DALTON: The molden file from dalton cannot be opened by jmol."o2_state_tpssd3_prod_oh_opt_cosmo.xyz" "o2_state_tpssd3_o_opt_cosmo.xyz"Įcho "load structure.xyz spacefill 0.2" > orbital_$i".spt"Įcho 'isosurface sign "'$i.cube'" translucent 0.4' > orbital_$i".spt"Įcho "write jpg $i.jpg" > orbital_$i".spt" "o2_state_tpssd3_react_oo_opt_cosmo.xyz" "o2_state_tpssd3_oh2_opt_cosmo.xyz" "o2_state_tpssd3_react_oh_opt_cosmo_singlet.xyz" "o2_state_tpssd3_react_ooh_opt_cosmo_singlet.xyz" Load FILES "o2_state_tpssd3_ooh2_opt_cosmo.xyz" The following java script can be inserted in the Jmol scripting window - It loads the files rotate them (adjust this according to your needs), writes the distances between atoms 49/50, 49/1, 49/10 and 49/30 and prints a frame in jpg format.Script to generate many MOs for i in do cubegen 0 MO="$i" orbitals.fchk "$i".cube done Jmol structures and molecular orbitals Note for an unrestricted calculation, use "AMO" or "BMO" instead. Use the utility program cubegen cubegen 0 MO= gaussian.fchk.Note: make sure the gaussian module is loaded (aurora: module load gaussian/16.B.01-AVX2) Transform the ".chk" with the utility program formchk gaussian.chk gaussian.fchk.Run a gaussian wave function optimization and make sure you obtain the ".chk" file, e.g.We focus here on generation of cube files that can be visualized with Jmol (see below). In gaussian, the can be generated in different ways and visualized with Jmol (see below). They are particularly useful for interpreting electronic spectroscopy (i.e. Molecular orbitals are often useful for understanding the electronic structure of a molecule. xyz or pdb file can be written out with from File => Save Coordinates.IMPORTANT: The align will happend according to the "top" molecule (marked with "T" in the VMD main window - this molecule will have unchanged coordinates) It only works if there are equally many of the atom types in the fwo files!). (note that they should have the same name in the pdb/xyz files that are compared. Now type for the atoms that should be aligned (in box upper left) name ATOM1 ATOM2 ATOM3 etc.Check that it is the right molecules that are read in (try "Add all"). Extensions => Analysis => RMSD Trajectory Tool.Note: this can also be done (often faster) with Maestro "Quick Align" Type vmd > set sel4 įor all examples, write the pdb using writepdb selection: vmd > $selX writepdb pdb_out.pdb.Select whole molecules, here waters - and only waters Save pdb with selection of atoms within 5 Å of another atom, here "Mn" type command set sel1 Ĭheck that the number of atoms fit by command $sel1numĮxample 2.Saving pdbs with different selections (examples) within 5 of name FE (selects all atoms withn 5 Å of FE). 
0 Comments
Leave a Reply. |
 RSS Feed
RSS Feed
